Mitochondria DNA Seen to Be Complex, Vary in Speed of Mutations Between People and Mice in Study
Mitochondria genetics may be more complex than previously thought, with new research reporting that DNA content can vary significantly among different mitochondria within a single cell, and between mice and people — a finding of potential importance to research that relies on mouse models of disease.
The work may also help to improve the understanding of diseases that arise from accumulated mutations in mitochondria, and provide new clues as to how individual patients may respond to therapies.
The study “Pervasive within-Mitochondrion Single-Nucleotide Variant Heteroplasmy as Revealed by Single-Mitochondrion Sequencing,” was published in Cell Reports.
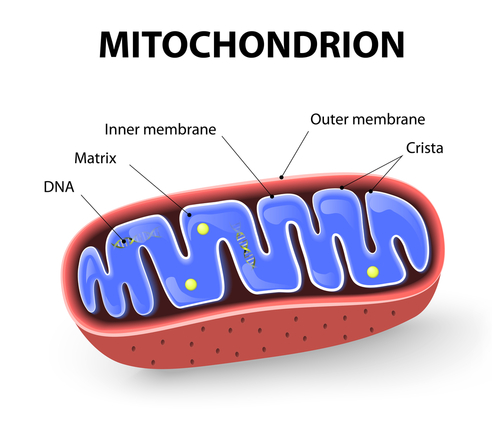
Each cell can have a few hundred to a few thousand mitochondria, each with their own DNA, called mtDNA. These organelles are responsible for producing all the energy that a cell needs to function and survive, and are also involved in other major cellular mechanisms like oxidative stress and calcium metabolism.
All the genetic information required to produce new mitochondria are encoded by its mtDNA, so that when mitochondrial DNA is affected by mutations, faulty mitochondria can be produced.
Mutations in mitochondrial genetic content have been linked to several diseases, including myoclonic epilepsy with ragged-red fiber disease (MERFF), mitochondrial encephalomyopathy, lactic acidosis, stroke-like symptoms (MELAS), and Leber’s hereditary optic neuropathy (LHON), as well as type 2 diabetes and cancer.
To better understand the genetics of mitochondria, a team of researchers at the University of Pennsylvania isolated single mitochondrion from cells and compared the mtDNA extracted from each.
Led by James Eberwine, a professor of Systems Pharmacology and Translational Therapeutics, the team extracted mitochondria from nerve cells collected from mice and people. They identified several genetic mutations in mtDNA sequences, and observed that mice mtDNA accumulated more mutations than human mtDNA.
This finding suggested that mutations in mtDNA may accumulate at a different pace in mice and in humans — a point with considerable implications on therapy discovery and development, since it is important to ensure that mice disease models closely relate to human disease.
“These results emphasize the need for human-based model systems for the study of mitochondrial genomics and disease,” the researchers wrote.
In addition, the differential genetic pattern found in isolated mitochondria may help in understanding the clinical variability seen in mitochondrial diseases and improve diagnosis. mtDNA mutation patterns could potentially be used to detect cells that could become diseased or to identify patients at risk of certain diseases.
“By being able to look at a single mitochondrion and compare mutational dynamics between mitochondria, we will be able to gauge the risk for reaching a threshold for diseases associated with increasing numbers of mitochondrial mutations,” Eberwine said in a university news release.
The team plan to use its mtDNA sequencing approach and knowledge gained to investigate ways of slowing the rate of accumulation in mtDNA mutations, and possibly mitochondrial disease progression.
“This roadmap of the location and number of mutations within the DNA of a mitochondrion and across all of a cell’s mitochondria is where we need to start,” Eberwine said.






